Scaling Vector Databases: Handle 100M+ Vectors Without Lag (2026)
A fintech startup built a fraud detection system on a single Qdrant node with 80 million transaction embeddings. It worked fine until Black Friday, when query volume spiked 40x. The node maxed out CPU trying to serve concurrent HNSW searches — HNSW traversal is single-threaded per query in most implementations. Latency went from 12ms to 1.8 seconds. The fraud detection system effectively went offline.
They could have sharded the index across four nodes before the event. Or added a query cache for repeated fraud pattern lookups. Or switched from HNSW to a batched GPU-accelerated search that handles concurrency better. None of these require changing the application code significantly — they are infrastructure and configuration changes.
Scaling vector databases is a distinct problem from scaling relational databases. The bottleneck is different, the sharding model is different, and the caching strategies are different. Getting this right before you need it is much easier than fixing it under load.
Concept Overview
Vector database scaling has three distinct bottlenecks, each requiring different solutions.
Memory — HNSW indexes must fit in RAM to deliver millisecond queries. 100M vectors at 1536 dims (float32) requires 600GB of RAM — before graph overhead. Memory is usually the first constraint hit at scale.
CPU/concurrency — HNSW graph traversal is CPU-intensive and does not parallelize across cores efficiently within a single query. Concurrent queries compete for CPU resources. A single powerful node often performs worse under high concurrency than multiple smaller nodes.
Index build time — Rebuilding an HNSW index over 100M vectors takes hours. Systems that need frequent full rebuilds (after model updates, data refreshes) must account for this downtime.
Solutions involve combinations of: horizontal sharding across nodes, quantization to fit more vectors in RAM, caching frequently repeated queries, read replicas to handle concurrency, and tiered storage for hot/cold vector separation.
How It Works
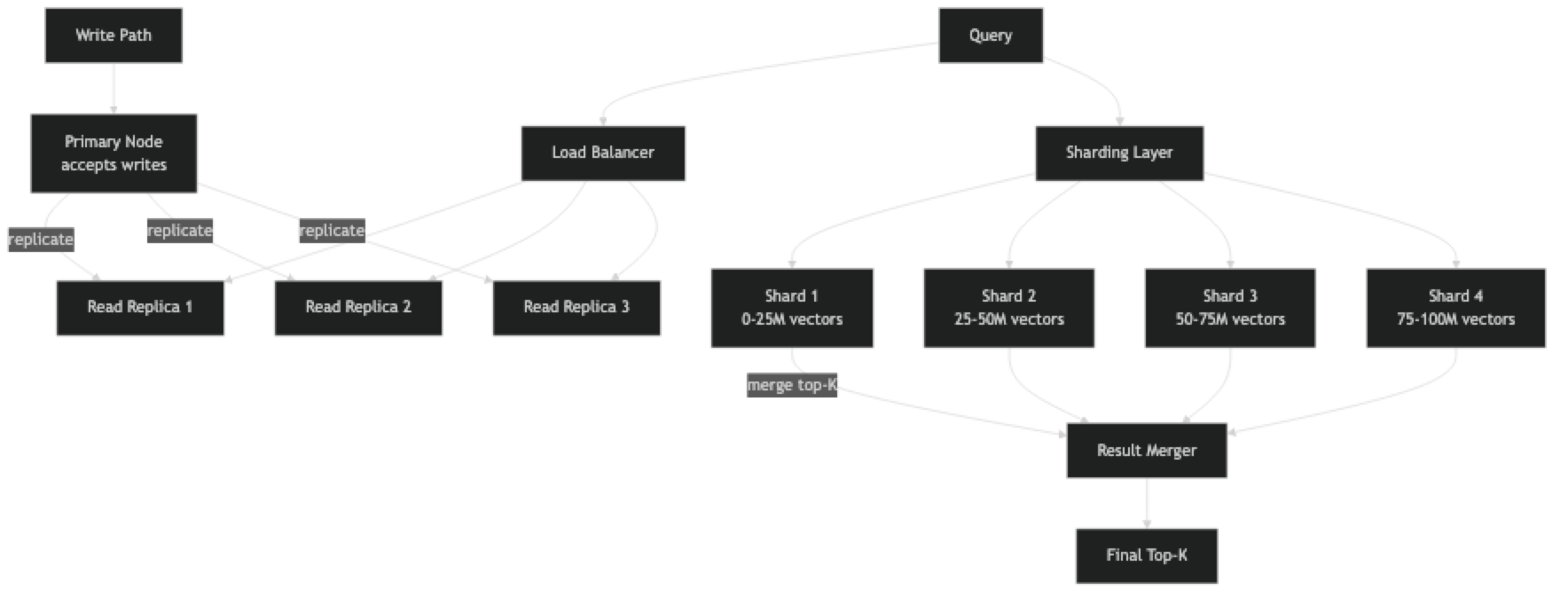
Sharding splits the vector corpus across multiple nodes. Each shard holds a fraction of the total vectors and its own HNSW index. At query time, the query is sent to all shards in parallel, each shard returns its local top-K, and a merge step selects the global top-K from the combined candidates.
This introduces a subtlety: to get a globally accurate top-10 across 4 shards, each shard must return more than 10 candidates (typically top-K × replication_factor candidates). The merge step then selects the best 10. The overhead is small — fetching 40 candidates from 4 shards and merging is trivially fast.
Replication (independent of sharding) runs multiple identical copies of each shard, distributing read queries across copies. This addresses the concurrency bottleneck: 4 replicas can handle 4x the concurrent query load.
Implementation Example
Query-Level Caching with Redis
Most vector search systems have a significant fraction of repeated or near-identical queries. For a RAG chatbot, "summarize this quarter's earnings" may be asked hundreds of times per day. Caching the embedding and result avoids redundant ANN searches.
pip install redis openai numpyimport redis
import hashlib
import json
import numpy as np
from openai import OpenAI
import chromadb
from chromadb.utils import embedding_functions
oai = OpenAI()
cache = redis.Redis(host="localhost", port=6379, decode_responses=False)
client = chromadb.PersistentClient(path="./prod_db")
ef = embedding_functions.OpenAIEmbeddingFunction(
api_key="sk-...", model_name="text-embedding-3-small"
)
collection = client.get_collection("documents", embedding_function=ef)
CACHE_TTL = 3600 # 1 hour — tune based on how often data changes
def cache_key(query: str, filters: dict, top_k: int) -> str:
payload = json.dumps({"q": query, "f": filters, "k": top_k}, sort_keys=True)
return f"vsearch:{hashlib.sha256(payload.encode()).hexdigest()}"
def search_with_cache(
query: str,
filters: dict | None = None,
top_k: int = 5,
) -> list[dict]:
key = cache_key(query, filters or {}, top_k)
# Check cache first
cached = cache.get(key)
if cached:
return json.loads(cached)
# Cache miss — query vector database
results = collection.query(
query_texts=[query],
n_results=top_k,
where=filters,
)
output = [
{"content": doc, "score": 1 - dist, "metadata": meta}
for doc, dist, meta in zip(
results["documents"][0],
results["distances"][0],
results["metadatas"][0],
)
]
# Store in cache
cache.setex(key, CACHE_TTL, json.dumps(output))
return output
results = search_with_cache("how does HNSW indexing work?", top_k=3)
for r in results:
print(f"[{r['score']:.3f}] {r['content']}")Sharded Search with Qdrant
pip install qdrant-clientfrom qdrant_client import QdrantClient
from qdrant_client.models import (
Distance, VectorParams, PointStruct,
Filter, FieldCondition, MatchValue,
)
import numpy as np
from concurrent.futures import ThreadPoolExecutor, as_completed
# Each shard is a separate Qdrant instance (or collection in multi-tenant setups)
SHARDS = [
QdrantClient(host="shard-1", port=6333),
QdrantClient(host="shard-2", port=6333),
QdrantClient(host="shard-3", port=6333),
QdrantClient(host="shard-4", port=6333),
]
COLLECTION = "documents"
DIM = 1536
def create_shards():
"""Initialize collection on each shard node."""
for shard in SHARDS:
shard.recreate_collection(
collection_name=COLLECTION,
vectors_config=VectorParams(size=DIM, distance=Distance.COSINE),
)
def shard_id_for_doc(doc_id: int) -> int:
"""Deterministic shard assignment by document ID."""
return doc_id % len(SHARDS)
def upsert_document(doc_id: int, vector: list[float], payload: dict) -> None:
shard_idx = shard_id_for_doc(doc_id)
SHARDS[shard_idx].upsert(
collection_name=COLLECTION,
points=[PointStruct(id=doc_id, vector=vector, payload=payload)],
)
def scatter_gather_search(
query_vector: list[float],
top_k: int = 10,
metadata_filter: dict | None = None,
fetch_k: int = 50, # fetch more per shard to get accurate global top-K
) -> list[dict]:
"""Query all shards in parallel, merge, return global top-K."""
qdrant_filter = None
if metadata_filter:
qdrant_filter = Filter(must=[
FieldCondition(key=k, match=MatchValue(value=v))
for k, v in metadata_filter.items()
])
def query_shard(shard: QdrantClient) -> list:
return shard.search(
collection_name=COLLECTION,
query_vector=query_vector,
limit=fetch_k,
query_filter=qdrant_filter,
)
# Parallel scatter
all_results = []
with ThreadPoolExecutor(max_workers=len(SHARDS)) as executor:
futures = {executor.submit(query_shard, shard): i for i, shard in enumerate(SHARDS)}
for future in as_completed(futures):
try:
results = future.result()
all_results.extend(results)
except Exception as e:
print(f"Shard {futures[future]} failed: {e}")
# Global merge — sort by score, take top-K
all_results.sort(key=lambda r: r.score, reverse=True)
return [{"id": r.id, "score": r.score, "payload": r.payload}
for r in all_results[:top_k]]
# Example usage
query_vec = np.random.randn(DIM).tolist()
results = scatter_gather_search(
query_vector=query_vec,
top_k=10,
metadata_filter={"category": "technical"},
)
for r in results:
print(f"[{r['score']:.3f}] id={r['id']} payload={r['payload']}")Embedding Cache to Reduce API Latency
Re-computing embeddings at query time adds 50–200ms of OpenAI API latency to every search. Caching query embeddings by hash eliminates this for repeated queries.
import redis
import hashlib
import numpy as np
from openai import OpenAI
oai = OpenAI()
cache = redis.Redis(host="localhost", port=6379, decode_responses=False)
def get_query_embedding(text: str, model: str = "text-embedding-3-small") -> np.ndarray:
key = f"emb:{model}:{hashlib.sha256(text.encode()).hexdigest()}"
cached = cache.get(key)
if cached:
return np.frombuffer(cached, dtype=np.float32)
response = oai.embeddings.create(model=model, input=text)
emb = np.array(response.data[0].embedding, dtype=np.float32)
cache.setex(key, 86400, emb.tobytes()) # cache for 24 hours
return embQuantization to Reduce Memory Footprint
import numpy as np
import faiss
DIM = 1536
N = 10_000_000 # 10M vectors
# Memory cost comparison
float32_gb = N * DIM * 4 / 1e9
int8_gb = N * DIM * 1 / 1e9 # scalar quantization: 4x reduction
pq_gb = N * 96 * 1 / 1e9 # product quantization with M=96: ~32x reduction
print(f"float32: {float32_gb:.1f} GB")
print(f"int8 SQ: {int8_gb:.1f} GB ({float32_gb/int8_gb:.0f}x reduction)")
print(f"PQ M=96: {pq_gb:.1f} GB ({float32_gb/pq_gb:.0f}x reduction)")
# Scalar quantization with FAISS
sq8 = faiss.IndexScalarQuantizer(DIM, faiss.ScalarQuantizer.QT_8bit, faiss.METRIC_INNER_PRODUCT)
# Or use HNSW with scalar quantization inside:
hnsw_sq = faiss.IndexHNSWSQ(DIM, faiss.ScalarQuantizer.QT_8bit, 32, faiss.METRIC_INNER_PRODUCT)
hnsw_sq.hnsw.efConstruction = 200Best Practices
Profile under realistic concurrency before scaling. Single-threaded benchmark numbers are misleading for HNSW. At 50 concurrent requests, a single HNSW node may be 10x slower than the single-thread benchmark suggests. Load test with realistic concurrency before choosing a scaling strategy.
Add read replicas before adding shards. Replication solves the concurrency bottleneck and is simpler than sharding. Sharding solves the dataset-too-large-for-one-node problem. If your dataset fits in one node's RAM, start with replicas only.
Use quantization to extend the single-node capacity before sharding. 4-bit or 8-bit quantization reduces memory by 4–8x with 2–5% recall loss. Going from 4 nodes to 1 node by quantizing is a large operational simplification.
Set appropriate efSearch per query tier. Not all queries have the same SLA. Internal analytics queries can afford efSearch=200 (higher recall, higher latency). User-facing autocomplete needs efSearch=50 (lower recall, lower latency). Most databases let you set this per-request.
Pre-warm indexes after node restarts. HNSW latency is 3–10x higher when the index is cold (not in OS page cache). After deploying or restarting a node, run a batch of warm-up queries before routing live traffic to it.
Common Mistakes
Sharding by random assignment when queries need geographic or tenant isolation. If your use case requires "search only documents owned by user X," sharding randomly across nodes means every query hits every shard. Shard by tenant or geographic region to avoid cross-shard fan-out for filtered queries.
Not accounting for index rebuild downtime. HNSW index rebuild over 100M vectors takes 4–8 hours on a single CPU node. If your embedding model updates quarterly, you need a strategy: build on a secondary node and swap, or maintain a continuous-update path. Pinecone's managed service handles this transparently; self-hosted systems do not.
Using synchronous embedding generation at index time for large datasets. Embedding 10M documents via OpenAI's API synchronously is a multi-day operation on a single thread. Use async batching with asyncio and the async OpenAI client — you can achieve 100–200 embeddings per second at scale.
Ignoring replication lag for write-heavy workloads. If you index new vectors and immediately query for them, replica lag means the new vectors may not appear in search results for seconds to minutes. Design your application to handle eventual consistency — do not assert that a newly indexed vector appears immediately in search.
Scaling the vector database without scaling the embedding service. If your system embeds queries at search time using a self-hosted model, that model becomes the bottleneck under high query load. Scale the embedding inference service (e.g., via TorchServe, vLLM, or Triton) in proportion to your vector database capacity.
Key Takeaways
- Vector database scaling has three distinct bottlenecks: memory (HNSW must fit in RAM), CPU concurrency (HNSW traversal does not parallelize well), and index rebuild time
- Add read replicas before adding shards — replication is simpler and solves the concurrency bottleneck without requiring scatter-gather query logic
- 8-bit scalar quantization reduces memory by 4x with ~2% recall loss — use it to extend single-node capacity before sharding
- Scatter-gather sharding requires each shard to return
top_k × replication_factorcandidates so the merge step can select the globally accurate top-K - Cache query embeddings in Redis to eliminate 50–200ms of embedding API latency for repeated queries
- Pre-warm HNSW indexes after every node restart — cold-start latency can be 3–10x higher than warm-cache latency
- Profile under realistic concurrent load before choosing a scaling strategy — single-thread benchmarks are misleading for HNSW
- Shard by tenant or data region, not randomly, when queries involve tenant-specific metadata filters
FAQ
How many vectors can a single node handle? With HNSW and float32 vectors, a node with 64GB RAM can hold roughly 10M vectors at 1536 dims (including graph overhead). With 8-bit quantization, this extends to ~40M vectors. With PQ compression, 100M+ vectors on a single node is feasible.
Does Pinecone handle sharding automatically? Yes. Pinecone's serverless tier automatically distributes data across their infrastructure. For pod-based Pinecone, you specify replicas (read concurrency) and pod type (performance tier) — sharding within a pod is internal. This is a primary reason teams choose Pinecone for production: you pay for managed scaling, not engineering time on it.
How do I handle index updates without downtime? The blue-green pattern: build the new index on a separate set of nodes while the old index serves traffic. Once the new index is validated (recall and latency benchmarks pass), switch the load balancer. Qdrant and Weaviate support live shard transfers that enable rolling updates without full cluster downtime.
What is the maximum recommended shard count for Qdrant or Weaviate? Qdrant recommends 1–3 shards per node and no more than 30 total shards per cluster for current versions. More shards increase the overhead of scatter-gather queries. The right shard count is determined by dataset size per node, not total cluster size — aim for 20–50GB of vector data per shard as a starting point.
How do I reduce HNSW index rebuild time for large datasets?
Parallelize insertion across multiple CPU cores (FAISS supports this with index.add() on multi-threaded builds). Use a GPU-accelerated build pipeline (FAISS has GPU indexes). Build on a secondary node while production traffic hits the current index, then swap. Reduce M (connections per node) to speed up build at the cost of slightly lower recall.
What is the right caching TTL for vector search results? TTL depends on how often your indexed data changes. For static knowledge bases (product catalogs, documentation), 1–24 hours is appropriate. For news or real-time data, 5–15 minutes. For user-specific results that personalize over time, do not cache results across users — only cache embeddings.
How do I handle hot spots in sharded vector search? Hot spots occur when one shard receives disproportionate query load (e.g., a popular category is mapped to one shard). Mitigation: add replicas to the hot shard, use content-based hash routing that distributes popular items across multiple shards, or use a managed service that handles load balancing automatically.