Vector Databases: Choose, Setup & Query in 30 Minutes (2026)
Your keyword search returns zero results for "ML model training cost" even though you have twelve articles on the topic. They use phrases like "compute budget," "GPU spend," and "training economics." A user searching with different vocabulary walks away with nothing.
This is the problem keyword-based search has never been able to solve — it matches tokens, not intent. Vector databases exist to fix exactly this. They store high-dimensional numerical representations of your data and let you search by meaning rather than lexical overlap.
In practice, almost every RAG system, semantic search engine, recommendation engine, and AI-powered feature you use today is backed by a vector database. Understanding how they work end-to-end — not just the API calls — determines whether your production system is fast, accurate, and cost-efficient, or none of those things.
Concept Overview
A vector database is a data store purpose-built for one operation: given a query vector, find the K stored vectors most similar to it.
To make that operation useful, you first convert your raw content (text, images, audio) into vectors using an embedding model. The embedding model maps similar inputs to nearby points in high-dimensional space. "Machine learning" and "deep learning" end up close together; "machine learning" and "sourdough bread" end up far apart.
The database then handles three concerns you do not want to manage yourself:
- Indexing — building a data structure that lets you search millions of vectors in milliseconds rather than seconds
- Persistence — storing both the vectors and associated metadata durably
- Filtering — combining vector similarity with structured predicates ("find the 10 most similar documents, but only from 2025")
One thing many developers overlook is that the embedding model and the vector database are separate concerns. You choose them independently, and a poor embedding model will produce bad results regardless of how well-tuned your database index is.
How It Works
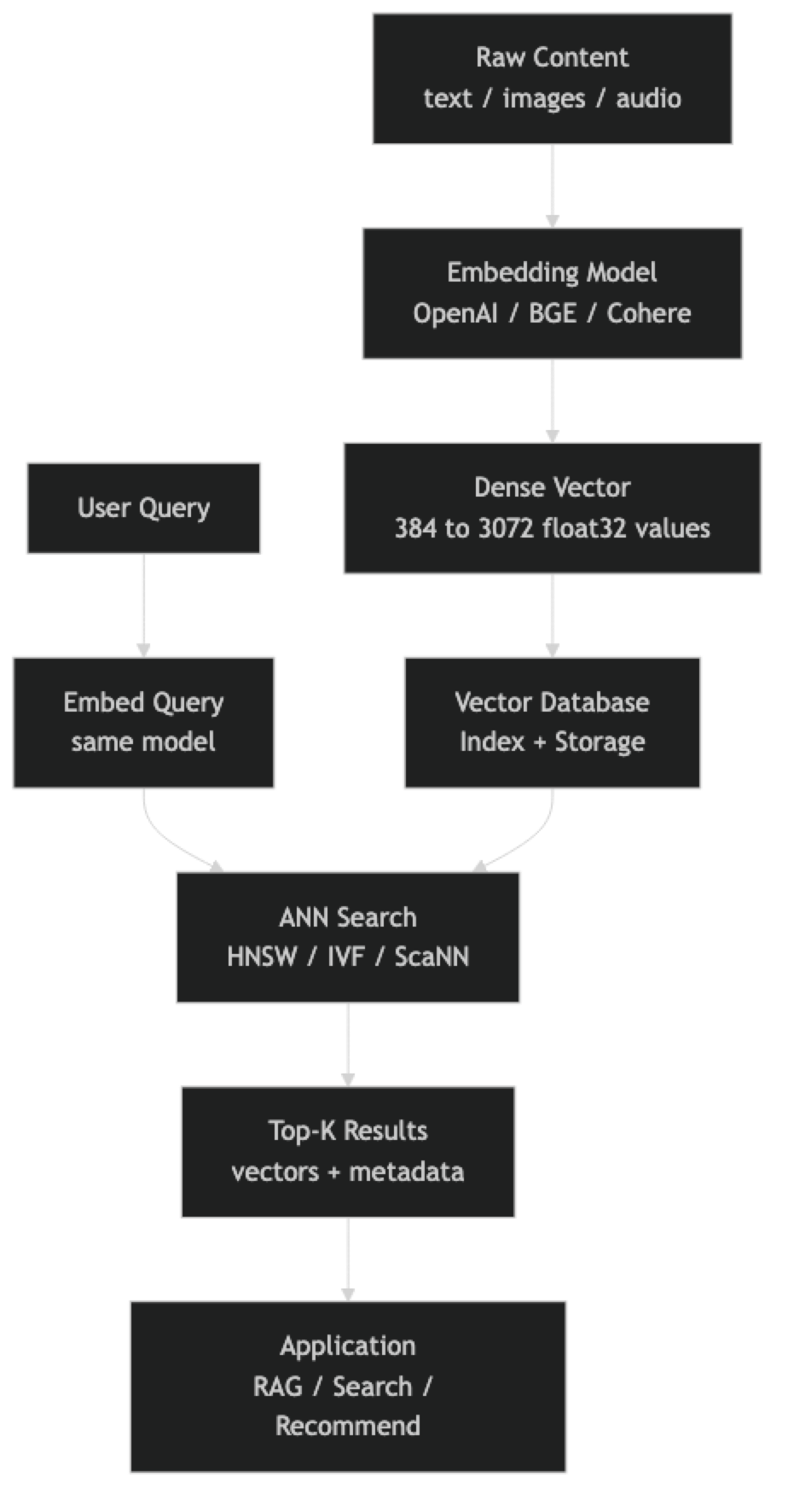
At index time, each piece of content goes through the embedding model to produce a float vector. That vector, along with its payload (original text, metadata), is inserted into the database. The database updates its index structure — most commonly HNSW — to incorporate the new vector.
At query time, the user's input is embedded with the same model. The database runs an approximate nearest neighbor search against the indexed vectors, returning the K most similar items by cosine similarity, dot product, or Euclidean distance.
The "approximate" in ANN is intentional. Exact nearest neighbor search requires comparing the query against every stored vector — O(n) complexity that stops scaling at roughly 100K vectors. ANN algorithms trade a small, tunable amount of recall for query times that stay in the single-digit milliseconds even at 100 million vectors.
Implementation Example
The fastest path to a working vector search system is ChromaDB — zero infrastructure, runs in-process, stores vectors on disk automatically.
pip install chromadb openaiimport chromadb
from chromadb.utils import embedding_functions
from openai import OpenAI
# Persistent local store — survives process restarts
client = chromadb.PersistentClient(path="./vector_store")
openai_ef = embedding_functions.OpenAIEmbeddingFunction(
api_key="sk-...",
model_name="text-embedding-3-small",
)
collection = client.get_or_create_collection(
name="engineering_docs",
embedding_function=openai_ef,
metadata={"hnsw:space": "cosine"},
)
# Index documents — Chroma handles embedding automatically
docs = [
"HNSW builds a multi-layer proximity graph for fast ANN search.",
"IVF partitions the vector space into Voronoi cells before searching.",
"Cosine similarity measures the angle between two vectors, ignoring magnitude.",
"Quantization compresses float32 vectors to int8 to reduce memory by 4x.",
"Metadata filtering lets you combine vector similarity with structured predicates.",
]
collection.add(
ids=[f"doc_{i}" for i in range(len(docs))],
documents=docs,
metadatas=[{"topic": "vector_db", "version": "2026"} for _ in docs],
)
# Query — returns results ordered by cosine similarity
results = collection.query(
query_texts=["How does approximate nearest neighbor search work?"],
n_results=3,
where={"topic": "vector_db"}, # optional metadata filter
)
for doc, distance in zip(results["documents"][0], results["distances"][0]):
print(f"[similarity: {1 - distance:.3f}] {doc}")For production workloads, Pinecone removes all infrastructure management:
pip install pinecone openaifrom pinecone import Pinecone, ServerlessSpec
from openai import OpenAI
import time
pc = Pinecone(api_key="your-api-key")
oai = OpenAI()
def embed(text: str) -> list[float]:
return oai.embeddings.create(
model="text-embedding-3-small",
input=text,
).data[0].embedding
# Create serverless index (us-east-1, free tier)
index_name = "engineering-docs"
if index_name not in [i.name for i in pc.list_indexes()]:
pc.create_index(
name=index_name,
dimension=1536,
metric="cosine",
spec=ServerlessSpec(cloud="aws", region="us-east-1"),
)
time.sleep(5) # wait for index to be ready
index = pc.Index(index_name)
# Upsert — batch for efficiency
records = [
("doc_0", embed("HNSW proximity graph for ANN"), {"topic": "indexing"}),
("doc_1", embed("IVF Voronoi cell partitioning"), {"topic": "indexing"}),
("doc_2", embed("cosine similarity for text embeddings"), {"topic": "metrics"}),
]
index.upsert(vectors=[
{"id": rid, "values": vec, "metadata": meta}
for rid, vec, meta in records
])
# Query with metadata filter
response = index.query(
vector=embed("How does vector indexing work?"),
top_k=5,
filter={"topic": {"$eq": "indexing"}},
include_metadata=True,
)
for match in response["matches"]:
print(f"[{match['score']:.3f}] id={match['id']} meta={match['metadata']}")For teams already running PostgreSQL, pgvector adds native vector search without a new service:
-- Install once per database
CREATE EXTENSION IF NOT EXISTS vector;
CREATE TABLE documents (
id SERIAL PRIMARY KEY,
content TEXT NOT NULL,
topic TEXT,
embedding vector(1536)
);
-- HNSW index for sub-millisecond search
CREATE INDEX ON documents
USING hnsw (embedding vector_cosine_ops)
WITH (m = 16, ef_construction = 64);
-- Insert
INSERT INTO documents (content, topic, embedding)
VALUES ('HNSW proximity graph', 'indexing', '[0.12, -0.34, ...]');
-- Search: <=> is cosine distance (lower = more similar)
SELECT content, topic,
1 - (embedding <=> '[0.12, -0.34, ...]') AS similarity
FROM documents
WHERE topic = 'indexing'
ORDER BY embedding <=> '[0.12, -0.34, ...]'
LIMIT 5;Best Practices
Batch your embedding calls. Every embedding model API accepts arrays. Embedding 1000 texts as one batch call is 20–50x faster than 1000 individual calls and costs the same.
Cache embeddings aggressively. If the same text will be embedded more than once — document chunks in a RAG system, product descriptions in a catalog — store the vector and reuse it. Re-embedding is pure waste.
Normalize vectors before storing. For cosine similarity, normalized vectors turn dot product into cosine similarity. Most databases do this automatically, but verifying saves debugging time.
Set ef_construction high at index build time. Building the HNSW index with ef_construction=200 produces a higher-quality graph than ef_construction=64. You only pay the build cost once. At query time, ef_search controls the speed/recall tradeoff dynamically.
Use metadata filtering to narrow candidate sets. Pre-filtering by date range, category, or user ID before running ANN search reduces both latency and the chance of irrelevant results surfacing. Every major vector DB supports this natively.
Monitor recall, not just latency. A fast search that consistently misses the best results is a broken search. Maintain a golden set of query/expected-result pairs and measure recall@10 as part of your deployment checks.
Common Mistakes
Using the wrong embedding model for the domain. Generic models trained on web text underperform on legal, medical, or code-heavy content. Always benchmark domain-specific alternatives — BAAI/bge-large-en-v1.5 or Cohere's specialized models — before locking in a choice.
Indexing full documents instead of chunks. Embedding a 10-page document into one vector dilutes the meaning. A query about section 3 can never surface that document reliably. Chunk at 256–512 tokens with 20% overlap before indexing.
Skipping the warm-up query. HNSW search on the first query after loading an index is often 5–10x slower than subsequent queries due to OS page cache cold starts. In production, run a few warm-up queries before serving traffic.
Confusing distance and similarity. Chroma returns cosine distance (lower is better). Pinecone and Qdrant return cosine similarity (higher is better). Sorting the wrong direction is a silent bug that produces terrible results.
Not setting a max_elements limit for in-memory indexes. Libraries like hnswlib require you to declare max capacity upfront. Exceeding it causes silent truncation or crashes. Always size for 2–3x your expected data volume.
Running brute-force search in production. If you have not explicitly created an index (HNSW, IVF, etc.), most databases fall back to brute-force O(n) search. Check your database's index status after data loads — pgvector is especially easy to misconfigure here.
Key Takeaways
- Vector databases store high-dimensional float vectors and retrieve the K most similar ones to a query vector using approximate nearest neighbor search
- The embedding model and the vector database are independent choices — a poor embedding model ruins results regardless of index quality
- HNSW is the default index for most vector databases and delivers single-digit millisecond query latency at millions of vectors
- Batch embedding API calls to reduce latency and cost by 20–50x compared to one-at-a-time calls
- Cache embeddings persistently — re-embedding the same content is pure waste with no quality benefit
- Always set
ef_constructionhigh at build time (200+) since you pay the cost once; tuneef_searchat runtime for the speed/recall tradeoff - Metadata filtering combined with vector similarity is the standard pattern for production RAG and search systems
- Start with ChromaDB locally, graduate to Pinecone or Qdrant for cloud production, and use pgvector if your team lives in PostgreSQL
FAQ
What is the difference between a vector database and a vector library like FAISS? A vector library (FAISS, hnswlib) is a pure indexing engine — it handles the ANN search math but nothing else. A vector database adds persistence, metadata storage, filtering, API access, and often replication. Use a library for embedded, single-process use cases; use a database for anything production-facing.
How many vectors can a vector database handle? It depends on the system. ChromaDB handles millions comfortably on a single machine. Pinecone and Weaviate have publicly demonstrated billion-vector scale with distributed architectures. pgvector with HNSW performs well to roughly 10–50 million vectors on a single PostgreSQL node before you need to consider sharding.
Can I use a vector database without an LLM? Yes. Vector databases are useful for any similarity search problem — product recommendation, image deduplication, music discovery, fraud pattern matching. LLMs are one consumer of vector search, not a requirement.
What embedding dimension should I use?
Higher dimensions capture more nuance but cost more memory and compute. text-embedding-3-small at 1536 dims is the right default for most English text use cases. Open-source models like BAAI/bge-small-en-v1.5 at 384 dims are a good choice when you need lower latency and cost.
Do I need to rebuild the index when I add new vectors? For HNSW-based databases (ChromaDB, Qdrant, Weaviate, Pinecone), no — the index is updated incrementally at insert time. For IVF-based indexes, adding significantly more vectors than the original training set degrades search quality and you should periodically rebuild.
How do I choose between ChromaDB, Pinecone, and pgvector? Use ChromaDB for local development and small-scale production where operational simplicity matters. Use Pinecone when you need managed cloud infrastructure that scales automatically without DevOps overhead. Use pgvector when your team already runs PostgreSQL and your dataset is under 5 million vectors.
What chunk size should I use when indexing documents? 256–512 tokens with 20% overlap is the standard starting point for most RAG and semantic search use cases. Shorter chunks lose context; longer chunks dilute the semantic signal. Adjust based on your document structure — technical documentation with focused sections does well at 128–256 tokens.