Vector Databases: Pinecone vs Qdrant vs Weaviate Tested (2026)
A team I know spent two weeks building a RAG prototype on FAISS. The semantic search worked well locally. Then they tried to add metadata filtering — "only search documents from the last 30 days" — and realized FAISS has no built-in metadata storage. They ended up maintaining a parallel SQLite database and joining results in Python. The whole approach collapsed under its own complexity.
Picking the wrong vector database does not just slow you down at the start — it creates architectural debt that compounds. The options differ significantly in what they actually are: some are indexing libraries, some are embedded databases, some are managed cloud services. Treating them as interchangeable is how you end up rebuilding your search layer at month three.
This comparison covers Chroma, Pinecone, FAISS, Weaviate, and pgvector with real code for each, so you can make an informed choice before committing.
Concept Overview
The vector database landscape splits into three categories with different deployment models and capabilities.
Indexing libraries (FAISS, hnswlib) — pure C++ search engines with Python bindings. They handle ANN math only. No persistence, no metadata, no API. Maximum performance, maximum manual work.
Embedded databases (ChromaDB, LanceDB) — run in-process alongside your application. No separate server. Good for development, single-node production, and applications where operational simplicity matters more than raw throughput.
Client-server / cloud databases (Pinecone, Weaviate, Qdrant, Milvus) — full database systems with REST APIs, replication, and scaling. Pinecone is fully managed SaaS. Weaviate, Qdrant, and Milvus can be self-hosted or cloud-managed.
One thing many developers overlook is that pgvector occupies a unique position — it adds vector search to an existing PostgreSQL instance, which means you get vector similarity inside a database that already handles your transactional data, ACID guarantees, and SQL joins.
How It Works
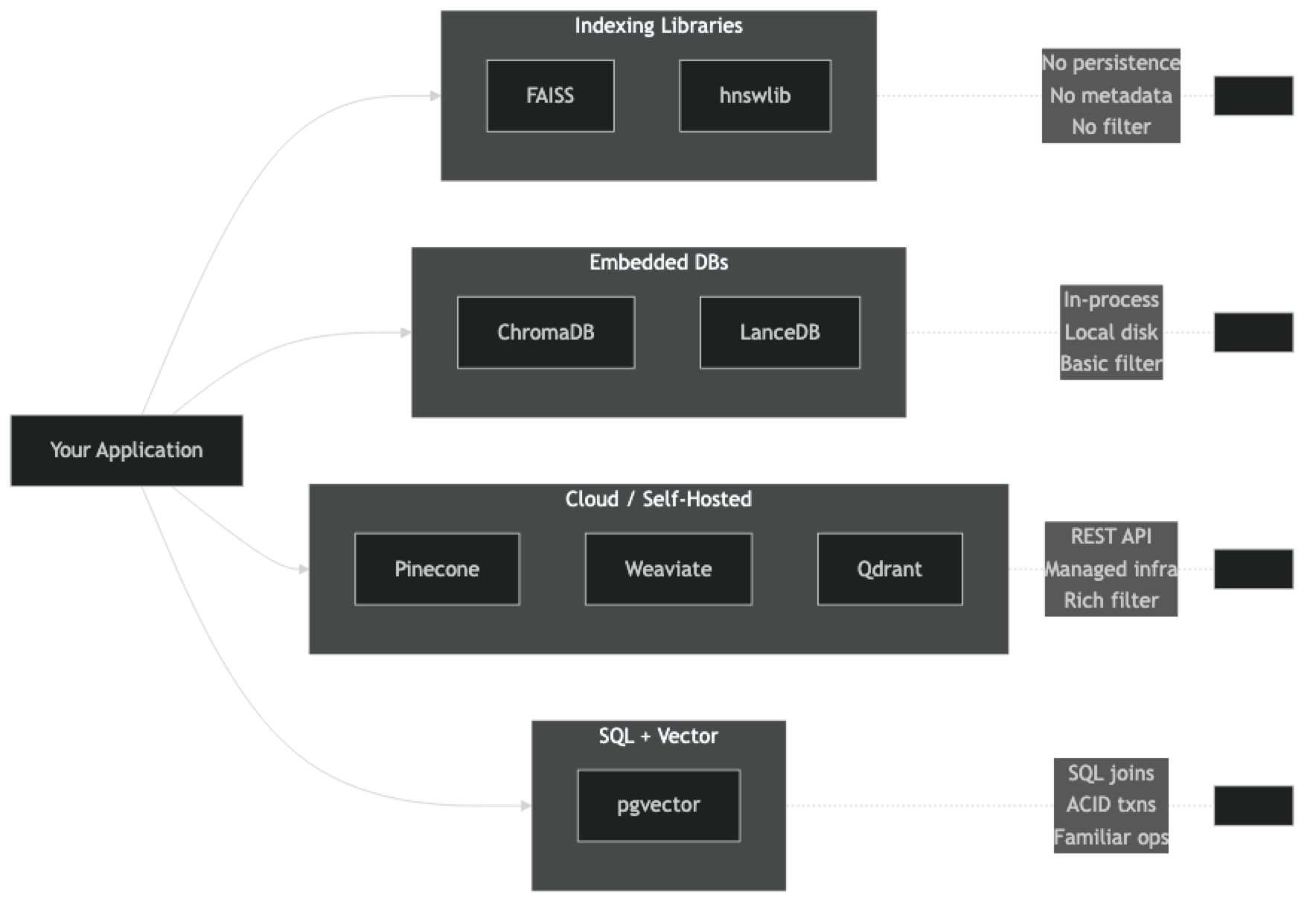
Each category makes a different tradeoff between operational complexity and feature richness. Libraries give you the lowest latency and the most control but require you to build everything around them. Cloud databases give you the most features out of the box but add a network round-trip to every query.
Implementation Example
ChromaDB — Local Development
pip install chromadb sentence-transformersimport chromadb
from chromadb.utils import embedding_functions
# Persistent client — data survives process restarts
client = chromadb.PersistentClient(path="./chroma_db")
# Use a free local embedding model
ef = embedding_functions.SentenceTransformerEmbeddingFunction(
model_name="BAAI/bge-small-en-v1.5"
)
collection = client.get_or_create_collection(
name="articles",
embedding_function=ef,
metadata={"hnsw:space": "cosine"},
)
collection.add(
ids=["a1", "a2", "a3", "a4"],
documents=[
"HNSW builds a hierarchical graph for fast approximate search.",
"IVF partitions vector space into clusters for efficient retrieval.",
"Cosine similarity measures the angle between two vectors.",
"Quantization reduces memory usage by compressing float32 to int8.",
],
metadatas=[
{"category": "indexing", "year": 2026},
{"category": "indexing", "year": 2026},
{"category": "metrics", "year": 2025},
{"category": "storage", "year": 2026},
],
)
# Query with metadata filter
results = collection.query(
query_texts=["approximate nearest neighbor algorithms"],
n_results=2,
where={"category": "indexing"},
)
for doc, dist in zip(results["documents"][0], results["distances"][0]):
print(f"[{1 - dist:.3f}] {doc}")FAISS — Maximum Performance
FAISS is the right tool when you need to embed ANN search inside a larger system and you will manage persistence yourself.
pip install faiss-cpu numpyimport faiss
import numpy as np
import pickle
DIM = 384 # match your embedding model's output dimension
# IndexFlatIP: exact search with inner product (use normalized vectors for cosine)
# IndexHNSWFlat: approximate search with HNSW graph
index = faiss.IndexHNSWFlat(DIM, 32) # 32 = M parameter
index.hnsw.efConstruction = 200
index.hnsw.efSearch = 50
# Add vectors (must be float32, C-contiguous)
vectors = np.random.randn(10_000, DIM).astype(np.float32)
faiss.normalize_L2(vectors) # normalize for cosine similarity
index.add(vectors)
# Search
query = np.random.randn(1, DIM).astype(np.float32)
faiss.normalize_L2(query)
distances, indices = index.search(query, k=5)
print(f"Top-5 indices: {indices[0]}")
print(f"Similarities: {distances[0]}") # dot product = cosine similarity after normalization
# Persist manually — FAISS has no built-in persistence
faiss.write_index(index, "hnsw.index")
with open("id_map.pkl", "wb") as f:
pickle.dump({"0": "doc_0", "1": "doc_1"}, f)
# Reload
index = faiss.read_index("hnsw.index")Pinecone — Managed Cloud
pip install pinecone openaifrom pinecone import Pinecone, ServerlessSpec
from openai import OpenAI
import time
pc = Pinecone(api_key="YOUR_PINECONE_KEY")
oai = OpenAI(api_key="YOUR_OPENAI_KEY")
def embed(texts: list[str]) -> list[list[float]]:
resp = oai.embeddings.create(model="text-embedding-3-small", input=texts)
return [r.embedding for r in resp.data]
# Create index once
if "articles" not in [i.name for i in pc.list_indexes()]:
pc.create_index(
name="articles",
dimension=1536,
metric="cosine",
spec=ServerlessSpec(cloud="aws", region="us-east-1"),
)
time.sleep(10)
index = pc.Index("articles")
# Upsert — batch for efficiency (max 100 per request)
texts = [
"HNSW hierarchical graph for ANN search",
"IVF clustering for vector retrieval",
"cosine similarity angle measurement",
]
vectors = embed(texts)
index.upsert(vectors=[
{"id": f"doc_{i}", "values": v, "metadata": {"text": t, "category": "indexing"}}
for i, (t, v) in enumerate(zip(texts, vectors))
])
# Query with metadata filter
result = index.query(
vector=embed(["approximate nearest neighbor"])[0],
top_k=3,
filter={"category": {"$eq": "indexing"}},
include_metadata=True,
)
for match in result["matches"]:
print(f"[{match['score']:.3f}] {match['metadata']['text']}")pgvector — SQL + Vectors
pip install psycopg2-binary pgvectorimport psycopg2
from pgvector.psycopg2 import register_vector
import numpy as np
conn = psycopg2.connect("postgresql://localhost/mydb")
register_vector(conn)
cur = conn.cursor()
# Setup (run once)
cur.execute("CREATE EXTENSION IF NOT EXISTS vector")
cur.execute("""
CREATE TABLE IF NOT EXISTS articles (
id SERIAL PRIMARY KEY,
content TEXT,
category TEXT,
embedding vector(1536)
)
""")
cur.execute("""
CREATE INDEX IF NOT EXISTS articles_embedding_idx
ON articles USING hnsw (embedding vector_cosine_ops)
WITH (m = 16, ef_construction = 64)
""")
conn.commit()
# Insert
from openai import OpenAI
oai = OpenAI()
def embed(text):
return oai.embeddings.create(model="text-embedding-3-small", input=text).data[0].embedding
content = "HNSW hierarchical graph for approximate search"
vec = embed(content)
cur.execute(
"INSERT INTO articles (content, category, embedding) VALUES (%s, %s, %s)",
(content, "indexing", vec)
)
conn.commit()
# Search — combine vector similarity with SQL WHERE
query_vec = embed("how does ANN indexing work?")
cur.execute("""
SELECT content, category,
1 - (embedding <=> %s) AS similarity
FROM articles
WHERE category = 'indexing'
ORDER BY embedding <=> %s
LIMIT 5
""", (query_vec, query_vec))
for row in cur.fetchall():
print(f"[{row[2]:.3f}] {row[0]} ({row[1]})")Best Practices
Match the database to your deployment model first. If you cannot run Docker in production, Weaviate and Qdrant self-hosted are off the table. If your company has a "no new vendor" policy, pgvector is your answer. Capability comparisons matter less than deployment constraints.
Always benchmark on your actual data and queries. Published benchmarks use synthetic data at specific dimensionalities. Your workload may hit different bottlenecks. Run a 10,000-vector pilot with real queries before committing.
Use serverless Pinecone for variable workloads. Serverless pricing scales to zero when idle and auto-scales under load. For RAG systems with unpredictable traffic, this is often cheaper than provisioning reserved capacity.
Prefer pgvector if you are already on PostgreSQL with under 5M vectors. You get vector search, metadata storage, ACID transactions, and SQL joins in one system. Adding a separate vector DB service is operational overhead you may not need.
Set FAISS's efSearch at query time, not at build time. You can tune the recall/latency tradeoff dynamically without rebuilding the index — useful for serving different SLA tiers from the same index.
Common Mistakes
Using FAISS in production without a metadata solution. FAISS stores vectors and integer IDs only. Every production use case needs metadata. Either use a database that includes it (ChromaDB, Pinecone, Qdrant) or maintain a parallel store and join — which quickly becomes painful.
Starting with Pinecone's starter pod and assuming performance scales linearly. Pinecone pod types (s1, p1, p2) have very different performance characteristics. Query latency at 10M vectors on a starter pod is not representative of production behavior. Test on the target pod type.
Not specifying the distance metric at collection creation time. ChromaDB and Pinecone both default to different metrics. If you change the metric after adding vectors, you must rebuild the collection — the indexes are incompatible. Specify metric="cosine" or hnsw:space=cosine explicitly.
Ignoring Weaviate's hybrid search capability. Weaviate supports BM25 + vector hybrid search natively with a single query. If your use case benefits from combining keyword and semantic signals (which most search use cases do), using Weaviate in pure-vector mode leaves significant quality on the table.
Key Takeaways
- FAISS is an indexing library, not a database — it has no persistence, metadata storage, or API, making it unsuitable for production use without significant wrapper work
- ChromaDB is the fastest path to a working local vector search system with zero infrastructure requirements
- Pinecone's serverless tier eliminates all operational overhead and scales to zero cost when idle — ideal for variable RAG workloads
- pgvector adds vector search directly to PostgreSQL and is the correct choice for teams already running PostgreSQL with under 5 million vectors
- Qdrant and Weaviate offer the best feature sets for self-hosted production: rich filtering, hybrid search, and horizontal scaling
- Always specify the distance metric at collection creation time — changing it later requires a full rebuild
- The most important decision criteria is deployment model compatibility, not benchmark numbers
- Run a benchmark on your actual data and query distribution before committing to any vector database
FAQ
Is FAISS a vector database? No — FAISS is a vector indexing library. It has no persistence, no metadata storage, and no query API. It is a building block that vector databases use internally. If you need a database, use ChromaDB, Qdrant, or Pinecone.
What is the cheapest vector database option? ChromaDB, Qdrant, and Weaviate are all open-source and free to self-host. The only cost is your infrastructure. Pinecone's serverless tier has a free quota sufficient for prototyping. pgvector adds no cost if you already run PostgreSQL.
Can I switch vector databases later without rebuilding my pipeline? You will need to re-embed your documents if you switch embedding models (not databases). Switching databases requires re-ingesting vectors but not re-embedding, assuming you kept the raw vectors. The main integration cost is the client library and API shape.
How does Weaviate's hybrid search work?
Weaviate runs BM25 keyword scoring and vector similarity scoring in parallel, then combines them with a weighted reciprocal rank fusion (RRF). The alpha parameter controls the balance: alpha=0 is pure BM25, alpha=1 is pure vector, alpha=0.5 blends both equally.
Which vector database has the lowest query latency? For local workloads, hnswlib and FAISS have the lowest latency because there is no network overhead. For cloud services, Pinecone's latency at p99 is typically under 50ms for indexes under 10M vectors. Qdrant self-hosted on a local GPU machine can achieve sub-millisecond p50 latency.
What is the difference between Qdrant and Weaviate? Both are open-source, self-hostable vector databases with cloud managed options. Qdrant is written in Rust and focuses on high-performance similarity search with rich payload filtering. Weaviate is written in Go and emphasizes hybrid search, a GraphQL API, and a module ecosystem for connecting embedding models. Choose Qdrant for raw performance; choose Weaviate if hybrid search and multi-modal capabilities matter.
How do I migrate from one vector database to another? Export your source documents and metadata (not necessarily the raw vectors). Re-insert into the new database using the same embedding model. If you cached raw embeddings separately, you can skip re-embedding. The main work is updating the client library calls — the query patterns are similar across all major vector databases.